VARIABILITY METHODS
USER GUIDE
VARIABILITY METHODS
Shannon Entropy
Shannon entropy analysis (Shannon, 1948 ) is possibly the most sensitive
tool to estimate the diversity
of a system.
For a multiple protein sequence alignment,
the Shannon entropy (H) for every position is as follow:
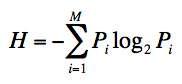
Where Pi is the fraction of residues of amino acid type
i, and M is the number of amino acid types (20).
H ranges from 0 (only one residue in present at that position) to 4.322
(all 20 residues are equally represented in that position). Typically,
positions with H >2.0 are considerered variable, whereas those with H < 2
are consider conserved. Highly conserved positions are those with H <1.0 (Litwin and Jores, 1992).
A minimum number of sequences is however required (~100) for H to describe the
diversity of a protein family.

Simpson Diversity Index
The Simpson index is another diversity index calculated from genotype proportions. This index describes the chance that two genotypes
sampled at random and with replacement from
a community will be from the same species. The value of this index ranges between 0 and 1,
the greater the value, the greater the sample diversity. Below is the
formula used to estimate it:
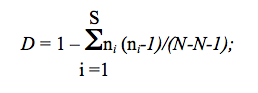
Where ni is the number of residues of type i, N is the total number of residues counted and S is number of different symbol types per site.

Wu-kabat Variability coefficient
The Wu-Kabat variability coefficient is a well-established descriptor of the susceptibility of
an amino acid position to evolutionary replacements(1977). It highlights stretches of accentuated
amino acid variation. The variability coefficient is computed using the following formula:
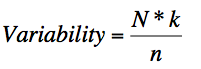
Where N is the number of sequences in the alignment, k is the number of different amino acids at a given position
and n is the frequency of the most common amino acid at that position.

USERGUIDE
Input
Protein Alignment
When this option is selected, a multiple sequence alignment (MSA) must be provided.
The alignment can either be pasted or uploaded from a file.
This program accepts the following MSA formats: Clustal,
FASTA and GCG/PileUp.
Only the standard 20 amino acids should be
included in the alignment. If other sequence characters are included
(e.g. X) the server will return an error message.
A typical example of Clustal alignment is the following:
CLUSTAL W (1.81) multiple sequence alignment
hla_a68w_1HSB SHSMRYFYTSVSRPGRGEPRFIAVGYVDDTQFVRFDSDAASQRMEPRAPWI
hla_a0201_1DUY SHSMRYFFTSVSRPGRGEPRFIAVGYVDDTQFVRFDSDAASQRMEPRAPWI
hla_b3501_1A1N SHSMRYFYTAMSRPGRGEPRFIAVGYVDDTQFVRFDSDAASPRTEPRPPWI
hla_b5301_1A1M SHSMRYFYTAMSRPGRGEPRFIAVGYVDDTQFVRFDSDAASPRTEPRPPWI
hla_b5101_1E27 SHSMRYFYTAMSRPGRGEPRFIAVGYVDDTQFVRFDSDAASPRTEPRAPWI
hla_b2701_1HSA SHSMRYFHTSVSRPGRGEPRFITVGYVDDTLFVRFDSDAASPREEPRAPWI
hla_cw3_1EFX SHSMRYFYTAVSRPGRGEPHFIAVGYVDDTQFVRFDSDAASPRGEPRAPWV
hla-cw4_1IM9 SHSMRYFSTSVSWPGRGEPRFIAVGYVDDTQFVRFDSDAASPRGEPREPWV
mkb_2vaa PHSLRYFVTAVSRPGLGEPRYMEVGYVDDTEFVRFDSDAENPRYEPRARWM
db-1BZ9 PHSMRYFETAVSRPGLEEPRYISVGYVDNKEFVRFDSDAENPRYEPRAPWM
.**:*** *::* ** **::: *****:. ******** . * *** *:
hla_a68w_1HSB RNTRNVKAQSQTDRVDLGTLRGYYNQSEAGSHTIQMMYGCDVGS
hla_a0201_1DUY GETRKVKAHSQTHRVDLGTLRGYYNQSEAGSHTVQRMYGCDVGS
hla_b3501_1A1N RNTQIFKTNTQTYRESLRNLRGYYNQSEAGSHIIQRMYGCDLGP
hla_b5301_1A1M RNTQIFKTNTQTYRENLRIALRYYNQSEAGSHIIQRMYGCDLGP
hla_b5101_1E27 RNTQIFKTNTQTYRENLRIALRYYNQSEAGSHTWQTMYGCDVGP
hla_b2701_1HSA RETQICKAKAQTDREDLRTLLRYYNQSEAGSHTLQNMYGCDVGP
hla_cw3_1EFX RETQKYKRQAQTDRVSLRNLRGYYNQSEAGSHIIQRMYGCDVGP
hla-cw4_1IM9 RETQKYKRQAQADRVNLRKLRGYYNQSEDGSHTLQRMFGCDLGP
mkb_2vaa RETQKAKGNEQSFRVDLRTLLGYYNQSKGGSHTIQVISGCEVGS
db-1BZ9 RETQKAKGQEQWFRVSLRNLLGYYNQSAGGSHTLQQMSGCDLGS
:*: * : * * .* ***** *** * : **::*.
The same multiple sequence alignment in FASTA Format:
>hla_b5101_1E27
SHSMRYFYTAMSRPGRGEPRFIAVGYVDDTQFVRFDSDAASPRTEPRAPWIRNTQIFKTN
TQTYRENLRIALRYYNQSEAGSHTWQTMYGCDVGP
>hla_a0201_1DUY
SHSMRYFFTSVSRPGRGEPRFIAVGYVDDTQFVRFDSDAASQRMEPRAPWIGETRKVKAH
SQTHRVDLGTLRGYYNQSEAGSHTVQRMYGCDVGS
>hla-cw4_1IM9
SHSMRYFSTSVSWPGRGEPRFIAVGYVDDTQFVRFDSDAASPRGEPREPWVRETQKYKRQ
AQADRVNLRKLRGYYNQSEDGSHTLQRMFGCDLGP
>mkb_2vaa
PHSLRYFVTAVSRPGLGEPRYMEVGYVDDTEFVRFDSDAENPRYEPRARWMRETQKAKGN
EQSFRVDLRTLLGYYNQSKGGSHTIQVISGCEVGS
>hla_b2701_1HSA
SHSMRYFHTSVSRPGRGEPRFITVGYVDDTLFVRFDSDAASPREEPRAPWIRETQICKAK
AQTDREDLRTLLRYYNQSEAGSHTLQNMYGCDVGP
>hla_cw3_1EFX
SHSMRYFYTAVSRPGRGEPHFIAVGYVDDTQFVRFDSDAASPRGEPRAPWVRETQKYKRQ
AQTDRVSLRNLRGYYNQSEAGSHIIQRMYGCDVGP
>hla_b3501_1A1N
SHSMRYFYTAMSRPGRGEPRFIAVGYVDDTQFVRFDSDAASPRTEPRPPWIRNTQIFKTN
TQTYRESLRNLRGYYNQSEAGSHIIQRMYGCDLGP
>db-1BZ9
PHSMRYFETAVSRPGLEEPRYISVGYVDNKEFVRFDSDAENPRYEPRAPWMRETQKAKGQ
EQWFRVSLRNLLGYYNQSAGGSHTLQQMSGCDLGS
>hla_a68w_1HSB
SHSMRYFYTSVSRPGRGEPRFIAVGYVDDTQFVRFDSDAASQRMEPRAPWIRNTRNVKAQi
SQTDRVDLGTLRGYYNQSEAGSHTIQMMYGCDVGS
An example of GCG/PileUp alignment would be:
PileUp
MSF: 95 Type: P Check: 5477 ..
Name: hla_a68w_1HSB oo Len: 95 Check: 5515 Weight: 0.0
Name: hla_a0201_1DUY oo Len: 95 Check: 4661 Weight: 10.0
Name: hla_b3501_1A1N oo Len: 95 Check: 4585 Weight: 10.0
Name: hla_b5301_1A1M oo Len: 95 Check: 4402 Weight: 10.0
Name: hla_b5101_1E27 oo Len: 95 Check: 4791 Weight: 10.0
Name: hla_b2701_1HSA oo Len: 95 Check: 3347 Weight: 10.0
Name: hla_cw3_1EFX oo Len: 95 Check: 4868 Weight: 10.0
Name: hla-cw4_1IM9 oo Len: 95 Check: 4736 Weight: 10.0
Name: mkb_2vaa oo Len: 95 Check: 4517 Weight: 10.0
Name: db-1BZ9 oo Len: 95 Check: 4055 Weight: 10.0
//
hla_a68w_1HSB SHSMRYFYTS VSRPGRGEPR FIAVGYVDDT QFVRFDSDAA SQRMEPRAPW
hla_a0201_1DUY SHSMRYFFTS VSRPGRGEPR FIAVGYVDDT QFVRFDSDAA SQRMEPRAPW
hla_b3501_1A1N SHSMRYFYTA MSRPGRGEPR FIAVGYVDDT QFVRFDSDAA SPRTEPRPPW
hla_b5301_1A1M SHSMRYFYTA MSRPGRGEPR FIAVGYVDDT QFVRFDSDAA SPRTEPRPPW
hla_b5101_1E27 SHSMRYFYTA MSRPGRGEPR FIAVGYVDDT QFVRFDSDAA SPRTEPRAPW
hla_b2701_1HSA SHSMRYFHTS VSRPGRGEPR FITVGYVDDT LFVRFDSDAA SPREEPRAPW
hla_cw3_1EFX SHSMRYFYTA VSRPGRGEPH FIAVGYVDDT QFVRFDSDAA SPRGEPRAPW
hla-cw4_1IM9 SHSMRYFSTS VSWPGRGEPR FIAVGYVDDT QFVRFDSDAA SPRGEPREPW
mkb_2vaa PHSLRYFVTA VSRPGLGEPR YMEVGYVDDT EFVRFDSDAE NPRYEPRARW
db-1BZ9 PHSMRYFETA VSRPGLEEPR YISVGYVDNK EFVRFDSDAE NPRYEPRAPW
hla_a68w_1HSB IRNTRNVKAQ SQTDRVDLGT LRGYYNQSEA GSHTIQMMYG CDVGS
hla_a0201_1DUY IGETRKVKAH SQTHRVDLGT LRGYYNQSEA GSHTVQRMYG CDVGS
hla_b3501_1A1N IRNTQIFKTN TQTYRESLRN LRGYYNQSEA GSHIIQRMYG CDLGP
hla_b5301_1A1M IRNTQIFKTN TQTYRENLRI ALRYYNQSEA GSHIIQRMYG CDLGP
hla_b5101_1E27 IRNTQIFKTN TQTYRENLRI ALRYYNQSEA GSHTWQTMYG CDVGP
hla_b2701_1HSA IRETQICKAK AQTDREDLRT LLRYYNQSEA GSHTLQNMYG CDVGP
hla_cw3_1EFX VRETQKYKRQ AQTDRVSLRN LRGYYNQSEA GSHIIQRMYG CDVGP
hla-cw4_1IM9 VRETQKYKRQ AQADRVNLRK LRGYYNQSED GSHTLQRMFG CDLGP
mkb_2vaa MRETQKAKGN EQSFRVDLRT LLGYYNQSKG GSHTIQVISG CEVGS
db-1BZ9 MRETQKAKGQ EQWFRVSLRN LLGYYNQSAG GSHTLQQMSG CDLGS
 PDB File
When this option is selected, a pdb file or a pdb code must be provided. A PDB file is a text file containing
the tridimensional (3D) coordinates of each atom in a protein. In addition, a PDB file may contain other information
on top of the coordinates abouth the protein and experimental settings used to obtain the 3D structure. PDBs are deposited in the
Brookhaven database under a 4 symbol code. By clicking
here you can see an example of a pdb file that corresponds to the structure of the biotinyl
domain of acetyl-coenzyme A.
PDB File
When this option is selected, a pdb file or a pdb code must be provided. A PDB file is a text file containing
the tridimensional (3D) coordinates of each atom in a protein. In addition, a PDB file may contain other information
on top of the coordinates abouth the protein and experimental settings used to obtain the 3D structure. PDBs are deposited in the
Brookhaven database under a 4 symbol code. By clicking
here you can see an example of a pdb file that corresponds to the structure of the biotinyl
domain of acetyl-coenzyme A.
For the pdb file input option, PVS will generate a multiple sequence
alignment from the sequence in the PDB file. Additionally, if a Chain identifier is given, the
program will select that chain from the PDB file. When no chain is provided, it will select the
first chain by default.
 Variability Methods
Variability Methods
Reference Sequence:
The reference sequence can either be a consensus sequence or the first sequence in the alignment.
The second choice is particularly useful if the user has additional information on a given sequence,
and wants to set it as the standard. By default, the consensus sequence is selected.

Output Tasks
PVS can perform several output task such as: Plot variability
, Mask variability in sequence , Return Conserved fragments
or Map structural variability.
By default, PVS will plot the sequence variability.
When Mask variability in sequence or Return Conserved fragments
are selected, a variability threshold
must be provided.
This parameter has to be set within the
range of 0 to 4.3 (default is 1.0) when Shannon is the selected variability method, and within the range of 0 to 1 (default is 0.46) when
Simpson is the selected variability method.
Those positions with a variability value above the selected threshold
are filtered out.Positions with a variability value under the selected threshold are considered of low variability (highly conserved).
If both Shannon and Simpson methods are selected, PVS will proceed considering the variability threshold as for Shannon.
Plot variability.
Plot variability consists of a graph of the sequence variability plotted against the selected
reference sequence in the alignment as shown below.However, if 'Map Structural Variability' has also been
selected, the sequence variability will be plotted against the sequence in the provided PDB file, when input is 'protein alignment'.
Please note that you must only select 'Plot Variability' as an output task if you wish have the sequence variability plotted
against the reference sequence in your protein alignment.
When several variability methods have been selected,
their graphs can be displayed by clicking on the method name

Mask variability in sequence
This option masks in the selected reference sequence those residues with a variability
greater or equal than the selected variability threshold. The variability
masked sequence is returned in FASTA format (Shown below). When the user clicks on the 'Run Epitope
Prediction' button, the returned FASTA sequence will be sent to the RANKPEP algorithm for the
anticipation of conserved T-cell epitopes.

Conserved Fragments
This option identifies those fragments (minimum length selected by user) in the selected reference
sequence consisting of consecutive residues whose variability is under the variability threshold.
These fragments are returned in a table sorted by their position in the sequence alignment.
Since sequence variability provides a means by which some pathogens escape the immune system,
this option and that of the sequence variability masking are relevant for vaccine design considerations.
It is important however to notice that relevant antigenic regions can be composed of conserved and
variant regions. Unfortunatelly, these fragments will not appear in the conserved fragments ouput
if they do not have the minimum number of consecutives conserved residues selected by the user.
Fragment Length:
This parameter sets the minimun length of the fragment. Each of the fragment
residues has a H that is under the threshold value .
Only the longest stretch of residues with H under the threshold is listed.

Map structural variability
This option maps the sequence variability onto a representative 3D-structure, using the PDB file
provided by the user. This is done using a JMOL applet, and for a correct visualization, javascript
must be enabled in the browser. By default, the 3-D structure is shown as 'wireframe', although other
display options can be selected by the user. For instance, in the image below, the selected option is
'trace'. The 'Back to original mapping' button will restore the sequence variability mapping when the
'Conserved Fragments' option has been selected and the user has clicked on a fragment to locate it
on the PDB file.

Shannon, C. E. (1948) The mathematical theory of communication.
The Bell system Technical Journal, 27, 379-423 & 623-656.
Kabat, E. A., Wu, T. T., and Bilofsky, H. (1977) Unusual distribution of amino acids
in complementarity-determing (hypervariable) segments of heavy and light chains
of immunoglobulins and their possible roles in specificity of antibody
combining sites.J. Biol. Chem.252, 6609-6616.
Litwin, S. and Jores, R. (1992) In theoretical and experimental insights
into immunology, (Edited by Perelson A. S. and Weisbuch G.), Springer-Verlag,
Berlin
Last change: November 2007
